library(tidyverse)
library(babynames)Data Opertations in the Tidyverse
Data Frame Recap
Filtering
babynames |>
filter(name == "Josef") |>
head()Summarising
babynames |>
summarise(
total = sum(n)
) |>
head()Filtering and Summarising
babynames |>
filter(
name == "Josef"
) |>
summarise(
total = sum(n)
) |>
head()Mutating
babynames |>
mutate(
name_length = nchar(name)
) |>
head()Split-Apply-Combine
What if we wanted a plot of how many total baby names there were each year?
We know how to get the total for 1 year with filtering and summarising.
babynames |>
filter(
year == 1880
) |>
summarise(
total = sum(n)
)But there a lot of unique years in the data
babynames |>
summarise(
min = min(year),
max = max(year)
) |>
mutate(
total_years = (max-min)+1
)I don’t want to filter and summarise 138 times.
.by or group_by()
babynames |>
summarise(
.by = year,
total = sum(n)
)- 1
- This will group the day by year, then apply the summarising within each group.
The .by argument is really handy. There’s another way to write it out that explicitly uses a function called group_by()
babynames |>
group_by(year) |>
summarise(
total = sum(n)
)- 1
- This will group the data frame, then any subsequent data verbs will apply within the grouping.
This goes one step further than what’s in the notes. Calculate the total number of baby names by year and sex.
To summarise just by year, we passed a single column name to summarise()
babynames |>
summarise(
.by = year,
total = sum(n)
)How do we construct a vector of values in R?
babynames |>
summarise(
.by = c(year, sex),
total = sum(n)
)More Split-Apply-Combine
Oakley and Skyler are collaborating on a programming project, and for some reason they want to keep track of how many lines of code they contribute on each day.
tibble(
day = c(
"Monday", "Tuesday",
"Monday", "Tuesday", "Wednesday"
),
name = c(
"Oakley", "Oakley",
"Skyler", "Skyler", "Skyler"
),
lines = c(
10, 30,
5, 5, 10
)
)->
collab_df
collab_df- 1
-
This will be the
daycolumn. - 2
-
This will be the
namecolumn. - 3
-
This will be the
linescolumn.
Oakley wants to know, on each day, what proportion of lines they wrote.
collab_df |>
mutate(
.by = day,
day_lines = sum(lines),
person_prop = lines/day_lines
)Skyler wants to know how their contributions are distributed across days.
This is still going to involve summing across the lines column.
collab_df |>
mutate(
.by = name,
total = sum(lines),
day_prop = lines/total
)Joining
Oakley and Skyler have, for some reason, wanted to include their year of birth in the data frame. So they create a separate yob data frame.
tibble(
name = c("Oakley", "Skyler"),
year = c(2000, 1995)
)->
yob
yobTo merge this yob data onto their colab_df data, they can use left_join().
collab_df |>
left_join(
yob
)->
collab_df
collab_dfOakley and Skyler are having an argument about which name was more popular in their respective years of birth. They’ve made a total_babies data frame like this:
babynames |>
summarise(
.by = c(year, name),
total_babies = sum(n)
)->
total_babies
total_babiesHow would they get the correct values for total_babies included as a column on collab_df?
Don’t overthink this one.
collab_df |>
left_join(total_babies)Practical Applications
Case 1: A vowel means plot
The Peterson & Barney dataset
library(phonTools)
data("pb52")
head(pb52)Let’s get the means for a plot
pb52 |>
summarise(
.by = c(type, repetition, vowel),
across(f0:f3, mean)
) ->
pb52_means
pb52_means- 1
-
This
across()function is handy way to apply the same operation (mean()) to a whole bunch of columns.
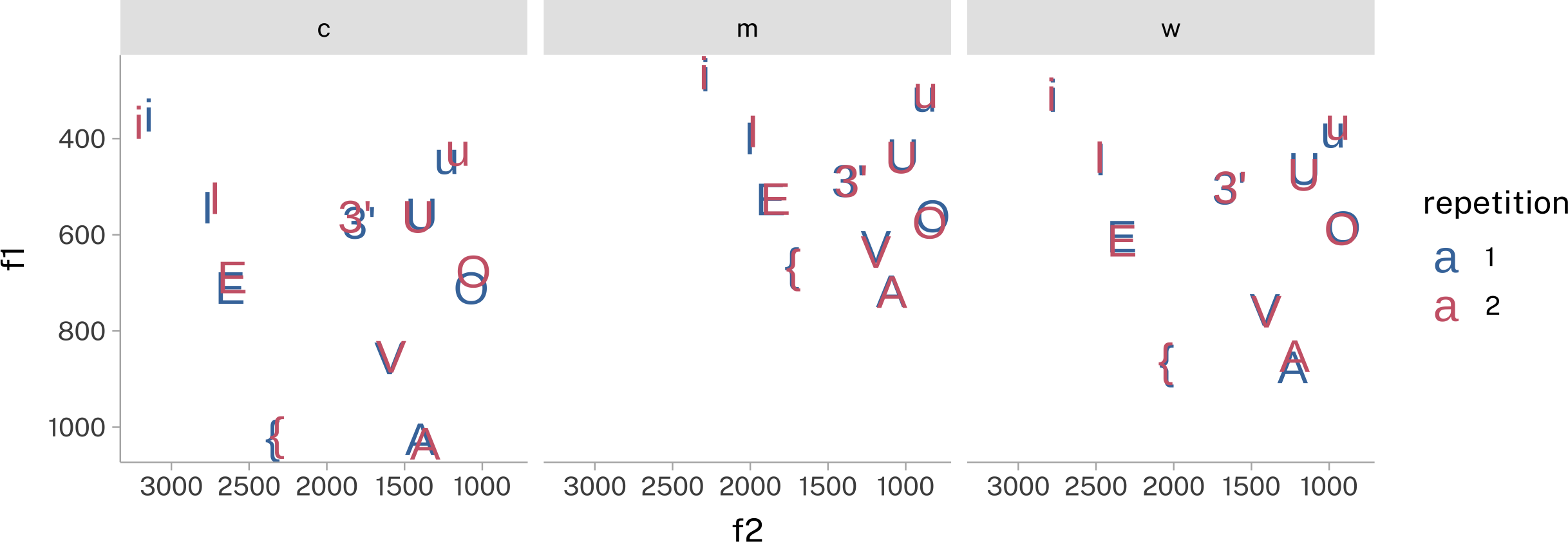
pb52_means |>
ggplot(
aes(
x = f2,
y = f1,
color = factor(repetition)
)
)+
geom_text(
aes(label=vowel),
size = 6
)+
scale_x_reverse()+
scale_y_reverse()+
facet_wrap(~type)+
labs(
color = "repetition"
)+
theme(
aspect.ratio = 1
)- 1
-
I have to change the data type of
repetitionso it gets categorical data coloring, not continuous coloring. - 2
- While not accurate to the data spans, for now I want the plotting area of each facet to be square. That is the height:width ratio = 1
But, that X-SAMPA isn’t everyone’s cup of tea. Here’s a translation data frame from X-SAMPA to IPA (although, there is a package called {ipa}which might be able to do this easier).
tibble(
vowel = c(
"{", "3'", "A",
"E", "i", "I",
"O", "u", "U",
"V"
),
ipa = c(
"æ", "ɝ", "ɑ",
"ɛ", "i", "ɪ",
"ɔ", "u", "ʊ",
"ʌ"
)
)->
vowel2ipa
vowel2ipa- 1
- These are the original labels in the data set.
- 2
- These are the new IPA labels I want to use, which I just copy-pasted from an internet keyboard.
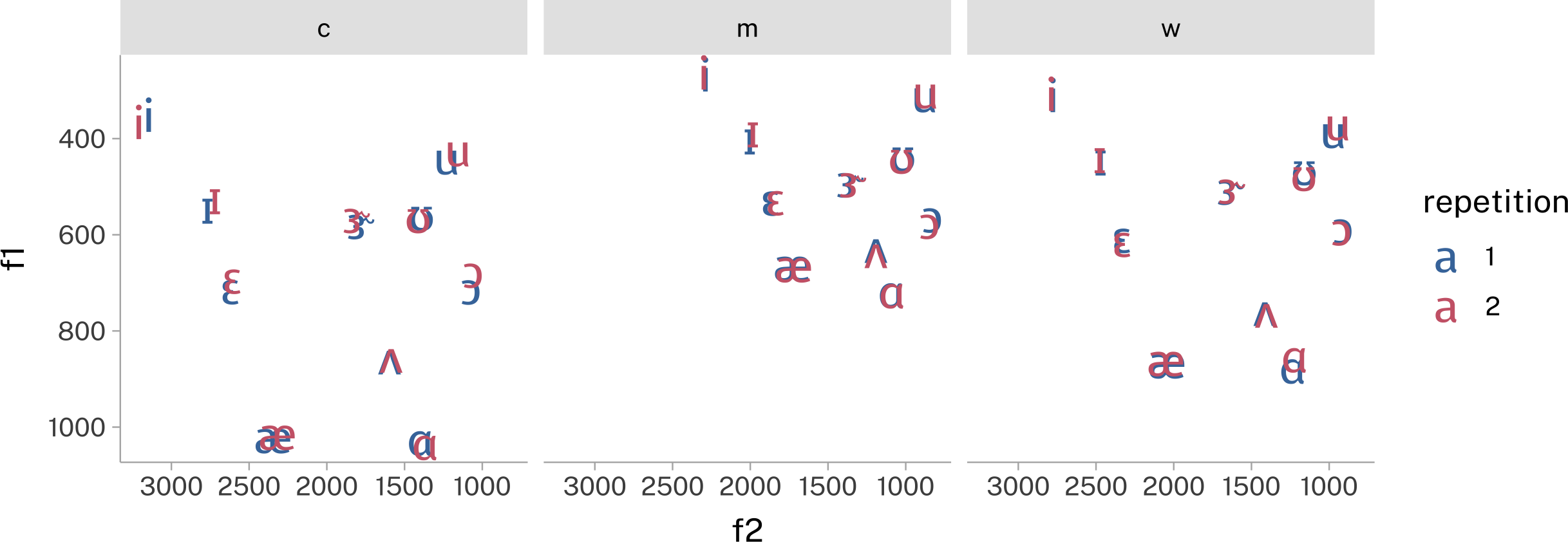
Now, you need to join the translation data frame onto the means data frame.
library(showtext)
font_add_google("Voces", "Voces")
showtext_auto()
pb52_means |>
left_join(vowel2ipa) |>
ggplot(
aes(
x = f2,
y = f1,
color = factor(repetition)
)
)+
geom_text(
aes(label=ipa),
size = 6,
family = "Voces"
)+
scale_x_reverse()+
scale_y_reverse()+
facet_wrap(~type)+
labs(
color = "repetition"
)+
theme(
aspect.ratio = 1
)- 1
- I needed to grab a font that could render the IPA symbols.
- 2
- The crucial `left_join()`.
- 3
- Setting the font family to be IPA ready.
Reuse
Citation
@online{fruehwald2024,
author = {Fruehwald, Josef},
title = {Data {Opertations} in the {Tidyverse}},
date = {2024-09-16},
url = {https://lin611-2024.github.io/notes/meetings/2024-09-16_data-operations.html},
langid = {en}
}